Last Updated: May 27, 2026
Introduction to the Congenital Disorders of Glycosylation
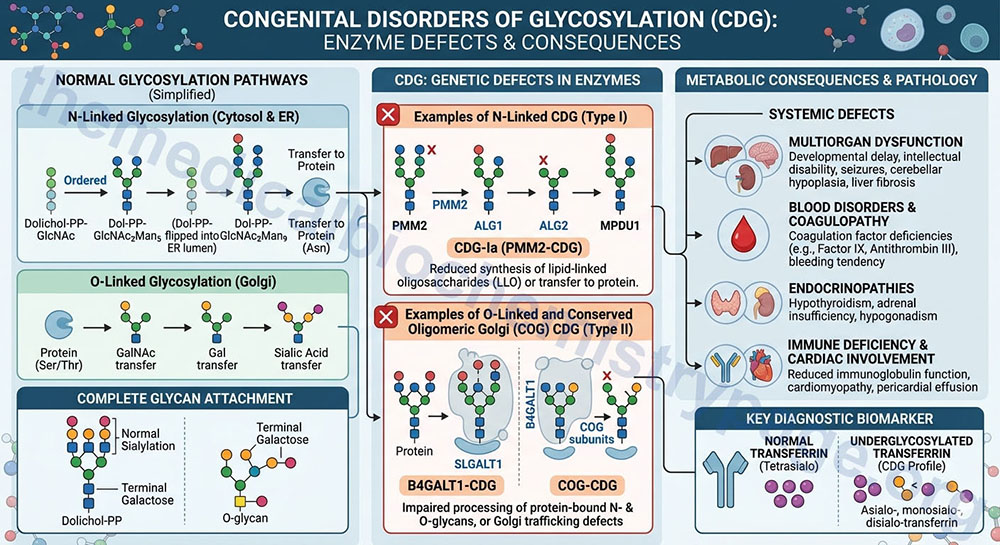
Congenital disorders of glycosylation (CDG) represent a constellation of diseases that result from defects in the synthesis of carbohydrate structures (glycans) and in the attachment of glycans to other compounds. At least 130 classified inherited disorders are known to be caused by mutations in genes encoding proteins involved in the synthesis of glycoconjugates with most of these disorders being of the CDG family.
The defective processes resulting in CDG involve the N-linked and O-linked glycosylation pathways, GPI-linkage, biosynthesis of proteoglycans as well as lipid glycosylation pathways. The majority of CDG are inherited as autosomal recessive disorders with few being autosomal dominant or X-linked. Given the range of disorders that result from CDG this page will focus primarily on the CDG that encompass defects in N-glycosylation of proteins. There are also syndromes related to defects in O-glycosylation and several are discussed in this page.
The CDG encompassing N-glycosylation defects are clustered into two broad categories. Group I CDG diseases are characterized by alterations/deficiencies in the synthesis and/or transfer of the Dol-PP-oligosaccharide precursor (termed the lipid linked oligosaccharide, LLO) to Asn residues in substrate proteins. Group II CDG diseases are characterized as those that result from defects in subsequent N-linked glycan processing. Current literature lists at least 28 CDG-I diseases (CDG-Ia – CDSG-Iz, MAGT1-CDG, and TUSC3-CDG) and at least 16 CDG-II diseases (CDG-IIa – CDG-IIp, ATP6V0A2-CDG). Proposed changes to the nomenclature of several CDG are indicated in parentheses following the current CDG identification. The disorder originally identified as CDG-Iv is actually a congenital disorder of deglycosylation (CDDG) resulting from mutations in the N-glycanase 1 gene (symbol: NGLY1).
Abbreviations used in the following descriptions include: Dol: dolichol; Fuc: fucose; Gal: galactose; Glc: glucose; GlcNAc: N-acetylglucosamine; Man: mannose; NANA: N-acetylneuraminic acid (sialic acid); P: phosphate
Type I CDG Forms
CDG-Ia (PMM2-CDG; CDG1A) is the most commonly occurring CDG, with appearance in individuals of European ancestry being highest. CDG-Ia is an autosomal recessive disease that results from mutations in the gene (PMM2) encoding phosphomannomutase 2 which is the enzyme that is required to convert Man-6-P to Man-1-P used in the generation of GDP-Man. Over 60 mutations in PMM2 have been identified that either decrease enzyme activity or stability.
The PMM2 gene is located on chromosome 16p13.2 and is composed of 8 exons that encode a 246 amino acid protein.
Although there is considerable variability in the clinical phenotypes observed in CDG-Ia patients, there is always some level of psychomotor impairment and in most cases it is quite profound. In addition, infants are ataxic, exhibit axial hypotonia, and have skeletal abnormalities consisting of long limbs and short torsos. Infants manifest with feeding problems that include vomiting, anorexia, and diarrhea. Other classical hallmarks of CDG-Ia are cerebellar hypoplasia and strabismus (lack of coordinated eye movements). Due to defective synthesis of coagulation factors by the liver (primarily factor XI, antithrombin III, protein C, and protein S), patients have severe coagulation defects. Adding to the situation is hepatomegaly with consequent liver dysfunction. About 20% of all CDG-Ia patients will die within the first year of life. However, most adults with the disorder have pleasant personalities even in the presence of stable intellectual impairment and variable peripheral neuropathy.
CDG-Ib (MPI-CDG; CDG1B) is an autosomal recessive disorder that results from defects in mannose phosphate isomerase (gene symbol: MPI) necessary to interconvert fructose-6-P and mannose-6-P. This enzyme carries out its function just prior to the role of PMM2 described for CDG-Ia. Deficiencies in MPI cause loss of plasma proteins via the intestines (protein-losing enteropathy, PLE), diarrhea and cyclical vomiting. Liver dysfunction is caused by hepatomegaly and results in coagulation defects and hypoglycemia. Thankfully, CDG-Ib patients are spared from intellectual defects. One explanation for the lack of demonstrable neurologic deficit in CDG-Ib compared to CDG-Ia is that brain hexokinase can phosphorylate mannose to Man-6-P thereby, bypassing the need for PMI. However, liver glucokinase does not phosphorylate mannose thus, there are the associated hepatic anomalies with CDG-Ib. Because free mannose is not found at any appreciable level in foods, dietary supplementation with mannose can ameliorate some of the symptoms of CDG-Ib.
The MPI gene is located on chromosome 15q24.1–q24.2 and is composed of 8 exons spanning 5kbp that generate five alternatively spliced mRNAs, each of which encode a distinct protein isoform.
CDG-Ic (ALG6-CDG; CDG1C) is an autosomal recessive disorder that results from defects in glucosyltransferase I (dolichylpyrophosphate Man9-GlcNAc2 α-1,3-glucosyltransferase; gene symbol: ALG6) The designation, ALG refers to Asparagine-Linked Glycosylation) and eight mutations have been identified in the ALG6 gene in patients with this disorder. ALG6 catalyzes the addition of the first glucose residue to the en bloc oligosaccharide that is subsequently N-linked to proteins. As a consequence of the defective ALG6 enzyme, the oligosaccharide unit is not transferred from dolichol to proteins. Symptoms of CDG-Ic are similar to those of CDG-Ia but much less severe. Patients have frequent seizures, psychomotor impairment that is milder than in CDG-Ia, pronounced axial hypotonia, and strabismus. Intestinal symptoms of CDG-Ic are markedly exacerbated by intestinal viral infections due to a decline of glycosylated structures (both proteins and proteoglycans) on the basolateral surfaces of intestinal enterocytes.
The ALG6 gene is located on chromosome 1p31.3 and is composed of 15 exons spanning 55kbp encoding a 507 amino acid precursor of a transmembrane protein.
Few cases of CDG-Id, CDG-Ie and CDG-If have been reported in the literature. Each of these three disorders are related via enzymes that synthesize or utilize Dol-P-Man structures.
CDG-Id (ALG3-CDG; CDG1E) is an autosomal recessive disorder that results from deficiencies in mannosyltransferase VI (Dol-P-Man: Man5GlcNAc2-P-P-Dol α-1,3-mannosyltransferase; gene symbol: ALG3). This enzyme transfers Man from Dol-P-Man to Dol-PP-Man5GlcNAc2 of the growing en bloc oligosaccharide. CDG-Id individuals suffer severe neurological impairment including profound psychomotor impairment and intractable seizures, eye abnormalities, optic atrophy, postnatal microcephaly, and hypsarrhythmia (abnormal EKG patterns observed in infants suffering spasms).
The ALG3 gene is located on chromosome 3q27.1 and is composed of 10 exons that generate at least two alternatively spliced mRNAs encoding ALG3 isoform a (438 amino acids) and ALG3 isoform d (390 amino acids).
CDG-Ie (DPM1-CDG; CDG1E) is an autosomal recessive disorder that results from defects in the catalytic subunit of Dol-P-Man synthase I (GDP-Man: Dol-P mannosyltransferase 1; gene symbol: DPM1). Dol-P-Man synthase I is an enzyme complex composed of at least three subunits identified as DPM1, DPM2, and DPM3. The catalytic subunit (DPM1) is stabilized via its association with the C-terminal domain of the DPM3 subunit and this interaction is stabilized by the DPM2 subunit. Dol-P-Man is required for mannose elongation reactions necessary to produce the en bloc oligosaccharide of N-glycans, GPI-anchored proteins and many other Man-containing glycoconjugates. Like patients with CDG-Id, CDG-Ie infants suffer severe neurological impairment including profound psychomotor impairment, intractable seizures, hypotonia, eye abnormalities, cortical blindness, and failure to thrive.
The DPM1 gene is located on chromosome 20q13.13 and is composed of 10 exons that generate four alternatively spliced mRNAs, each of which encode a distinct protein isoform.
CDG-If (MPDU1-CDG; CDG1F) is an autosomal recessive disorder that results from defects in the protein responsible for utilization of Dol-P-Man, independent of DPM1 which is defective in CDG-Ie. The gene encoding this activity is identified as Man-P-Dol utilization defect 1 (gene symbol: MPDU1) and it is required for the utilization of Dol-P-Man and Dol-P-Glc. The MPDU1 gene was identified on the basis of the fact that it could rescue defects in Chinese hamster ovary (CHO) cells that prevented the synthesis and utilization of Dol-P-Man. The CHO cell mutations were identified as Lec15 and Lec35 (synthesis and utilization, respectively) and the rescue gene was called SL15 for suppressor of Lec15. This gene was shown to rescue both the Lec15 and Lec35 defects.
Persons with CDG-If have clinical symptoms including dwarfism, hypotonia, frequent seizures, severe psychomotor impairment and cerebral atrophy. Additionally, these individuals have severe failure to thrive after birth.
The MPDU1 gene is located on chromosome 17p13.1 and is composed of 7 exons that generate two alternatively spliced mRNAs that encode a 247 amino acid transmembrane protein and a 192 amino acid protein. Both the N-terminus and C-terminus of the 247 amino acid MPDU1 protein are localized to the cytoplasm.
CDG-Ig (ALG12-CDG; CDG1G) is an autosomal recessive disorder that results from deficiencies in mannosyltransferase VIII (Dol-P-Man: Man7-GlcNAc2-P-P-Dol α-1,6-mannosyltransferase; gene symbol: ALG12). The common clinical features associated with CDG-Ig are psychomotor impairment, facial dystrophism, and hypotonia. In some patients there are feeding problems, microcephaly, convulsions, and frequent respiratory tract infections.
The ALG12 gene is located on chromosome 22q13.33 and is composed of 12 exons that encode a protein of 488 amino acids.
CDG-Ih (ALG8-CDG; CDG1H) is an autosomal recessive disorder that results from deficiencies in glucosyltransferase II (Dol-P-Glc: Glc1-Man9-GlcNAc2-P-P-Dol α-1,3-glucosyltransferase; gene symbol: ALG8). To date five children have been identified with CDG-Ih. Their symptoms included moderate hepatomegaly, PLE (see CDG-Ib), hypotonia, lung hypoplasia, anemia, and thrombocytopenia. All but one of the observed patients succumbed the the disorder within the first year of life.
The ALG8 gene is located on chromosome 11q14.1 and is composed of 14 exons that generate two alternatively spliced mRNAs encoding isoform a (526 amino acids) and isoform b (467 amino acids) versions of the enzyme. The ALG8 isoform b protein has not been conclusively demonstrated to possess the glucosyltransferase activity of the ALG8 isoform a protein.
CDG-Ii (ALG2-CDG; CDG1I) is an autosomal recessive disorder that results from deficiencies in mannosyltransferase II (GDP-Man: Man1-GlcNAc2-P-P-Dol α-1,3/1,6-mannosyltransferase; gene symbol: ALG2). The single patient thus far identified with this form of CDG exhibited severe psychomotor impairment, spasms, hypsarrhythmia, and irregular nystagmus (involuntary eye movements). Examination of the brain by MRI showed significant dysmyelination.
The ALG2 gene is located on chromosome 9q22.33 and is composed of 3 exons that encode a protein of 416 amino acids.
CDG-Ij (DPAGT1-CDG; CDG1J) is an autosomal recessive disorder that results from deficiencies in UDP-N-acetylglucosamine-dolichylphosphate N-acetylglucosaminephosphotransferase (gene symbol: DPAGT1). A single patient has been described with this form of CDG. The infant suffered from intractable spasms, was hypotonic and had sever psychomotor impairment. Dysmorphia was noted with microcephaly, micrognathia (undersized jaw), and estropia (a form of strabismus where one or both eyes turn inward).
The DPAGT1 gene is located on chromosome 11q23.3 and is composed of 9 exons that encode a protein of 408 amino acids.
CDG-Ik (ALG1-CDG; CDG1K) is an autosomal recessive disorder that results from deficiencies in mannosyltransferase I, also called chitobiosyldiphosphodolichol β-mannosyltransferase (GDP-Man: GlcNAc2-P-P-Dol β-1,4-mannosyltransferase; gene symbol: ALG1). Each of the infants that were identified with this form of CDG exhibited seizures, severe psychomotor impairment, and early death. Variable symptoms in the patients included hypotonia, dysmorphia, microcephaly, liver dysfunction, severe infections, and coagulation defects.
The ALG1 gene is located on chromosome 16p13.3 and is composed of 14 exons that generate two alternatively spliced mRNAs that encode proteins of 464 amino acids (isoform 1 ) and 353 amino acids (isoform 2).
CDG-IL (ALG9-CDG; CDG1L) is an autosomal recessive disorder that results from deficiencies in mannosyltransferase VII-IX (Dol-P-Man: Man6– and Man8-GlcNAc2-P-P-Dol α-1,2-mannosyltransferase; gene symbol: ALG9). Two patients with CDG-IL have been identified. Both exhibited psychomotor impairment, hypotonia, hepatomegaly, microcephaly, and seizures.
The ALG9 gene is located on chromosome 11q23.1 and is composed of 27 exons that generate 19 alternatively spliced mRNAs that collectively encode 12 different protein isoforms.
CDG-Im (DOLK-CDG; CDG1M) is an autosomal recessive disorder that results from deficiencies in the dolichol kinase gene. Dolichol kinase is the enzyme responsible for the terminal step in the synthesis of dolichol phosphate. At least 17 children have been identified with CDG-Im. The range of symptoms and age of presentation was highly variable in the identified patients. Symptoms include dilated cardiomyopathy, hypotonia, minimal hair growth, seizures, ichthyosis, and hypoketotic hypoglycemia. All the patients succumbed to cardiac failure at varying ages from 8 months to 13 years.
The DOLK gene is located on chromosome 9q34.11 and is an intronless gene that encodes a protein of 538 amino acids.
CDG-In (RFT1-CDG; CDG1N) is an autosomal recessive disorder that results from deficiencies in RFT1 homolog gene. The RFT1 homolog [human homolog of the yeast Rft1 (Requiring fifty-three protein 1) enzyme] is a flippase enzyme that is responsible for the translocation of the Man5-GlcNAc2-P-P-Dol intermediate from the cytoplasmic side to the luminal side of the endoplasmic reticulum (ER) membrane. Eight patients with CDG-In have been identified all of whom manifested symptoms in early childhood that included marked developmental delay, hypotonia, early-onset seizures, hepatomegaly, sensorineural deafness, and coagulopathy.
The RFT1 gene is located on chromosome 3p21.1 and is composed of 19 exons that encode a protein of 541 amino acids.
CDG-Io (DPM3-CDG; CDG1O) is an autosomal recessive disorder that results from deficiencies in dolichol-phosphate mannosyltransferase subunit 3 (gene symbol: DPM3). Dolichol-phosphate mannose (Dol-P-Man) is synthesized on the cytosolic side of the endoplasmic reticulum (ER) membrane from GDP-Man and dolichol-phosphate by the action of the dolichol-phosphate mannosyltransferase (DPM) complex of which the DPM3 encoded protein is a subunit. To date a single patient with CDG-Io has been identified who presented at age 11 with symptoms of dilated cardiomyopathy, muscle weakness, and an unsteady gait. The activity of the DPM complex is also required for the O-mannosylation of α-dystroglycan of the dystrophin glycoprotein complex (DGC) which links the congenital disorders of glycosylation to the dystroglycanopathies.
The DPM3 gene is located on chromosome 1q22 and is composed of 2 exons that generate two alternatively spliced mRNAs encoding an isoform 1 protein (122 amino acids) and an isoform 2 protein (92 amino acids).
CDG-Ip (ALG11-CDG; CDG1P) is an autosomal recessive disorder that results from deficiencies in GDP-Man:Man3GlcNAc2-PP-dolichol-α1,2-mannosyltransferase (gene symbol: ALG11). Five patients with CDG-Ip have been identified. All five infants exhibited severely delayed psychomotor development, intellectual impairment, seizures, hypotonia, dysmorphic features, and early lethality.
The ALG11 gene is located on chromosome 13q14.3 and is composed of 4 exons that encode a 492 amino acid protein.
CDG-Iq (SRD5A3-CDG; CDG1Q) is an autosomal recessive disorder that results from deficiencies in steroid 5α-reductase 3 (gene symbol: SRD5A3). The SRD5A3 gene is one of three related genes encoding steroid 5α-reductase enzymes that convert testosterone to dihydrotestosterone, DHT. In addition to DHT formation, the SRD5A3 encoded enzyme (also called polyprenol reductase) is required for the conversion of polyprenol to dolichol which is necessary for the synthesis of N-linked glycoproteins. Symptoms observed in patients harboring mutations in the SRD5A3 gene include ocular colobomas, ichthyosis, endocrine abnormalities associated with midline brain malformations, intellectual impairment, hypotonia, motor delay, and facial dysmorphism.
The SRD5A3 gene is located on chromosome 4q12 and is composed of 6 exons that encode a protein of 318 amino acids.
CDG-Ir (DDOST-CDG; CDG1R) is an extremely rare autosomal recessive disorder that results from deficiencies in dolichyl-diphosphooligosaccharide-protein glycosyltransferase non-catalytic subunit (gene symbol: DDOST). The DDOST encoded protein is a component of the N-oligosaccharyltransferase complex (OST) which is the enzyme complex that transfers lipid-linked oligosaccharide (LLO) structure (Glc3Man9GlcNAc2-Dol) to the asparagine residue in nascent polypeptides. This reaction is the central step in N-linked glycosylation of proteins. The initial identification of DDOST-CDG was a patient who presented with failure to thrive, gastroesophageal reflux, developmental delay, ear infections, oromotor dysfunction (defects in sensory inputs, motor systems, and movement organization that involves sucking, chewing, swallowing, speech articulation, and facial non-verbal communication), hypotonia, external strabismus, mild to moderate liver dysfunction, and delayed psychomotor development. A total of only three patients worldwide have been identified with DDOST-CDG.
The DDOST gene is located on chromosome 1p36.12 and is composed of 11 exons that encode a 456 amino acid precursor protein.
CDG-Is (ALG13-CDG; CDG1S) is an X-linked recessive disorder that results from deficiencies in a subunit of the heteromeric UDP-N-acetylglucosamine transferase (gene symbol: ALG13). Six patients with CDG-Is have been identified each with symptoms of early infantile epileptic encephalopathy. The symptoms in these infants were identified as early infantile epileptic encephalopathy-36 (EIEE36) and they are associated with defective protein glycosylation.
The ALG13 gene is located on the X chromosome (Xq23) and is composed of 35 exons that generate 17 alternatively spliced mRNAs that encode 11 different protein isoforms.
CDG-It (PGM1-CDG; CDG1T) is an autosomal recessive disorder that results from deficiencies in phosphoglucomutase 1 (gene symbol: PGM1). This congenital disorder of glycosylation was also known as glycogen storage disease type 14 (GSD14). Patients with this form of CDG manifest with cleft lip, bifid uvula, short stature, hepatomegaly, hypoglycemia, exercise intolerance.
The PGM1 gene is located on chromosome 1p31.3 and is composed of 13 exons that generate three alternatively spliced mRNAs each of which encodes a distinct protein isoform.
CDG-Iu (DPM2-CDG; CDG1U) is an autosomal recessive disorder that results from deficiencies in dolicholphosphate mannosyltransferase 2, regulatory subunit encoded by the DPM2 gene. DPM2-CDG is commonly called congenital muscular dystrophy with intellectual disability and severe epilepsy. Three patients from two unrelated families have been reported with CDG-Iu. All three exhibited psychomotor impairment, hypotonia, hepatomegaly, microcephaly, seizures, and early lethality.
The DPM2 gene is located on chromosome 9q34.11 and is composed of 5 exons that generate three alternatively spliced mRNAs, each of which encode a distinct protein isoform.
MAGT1-CDG is an X-linked recessive disorder that results from deficiencies in the magnesium transporter 1 gene (MAGT1). The magnesium transporter 1 protein interacts with N-oligosaccharyltransferase complex (OST) which is the enzyme complex that transfers the lipid-linked oligosaccharide (LLO) structure (Glc3Man9GlcNAc2-Dol) to the asparagine residue in nascent polypeptides. This reaction is the central step in N-linked glycosylation of proteins. The interaction of the MAGT1 protein with the OST indicates the protein has a role in protein N-glycosylation accounting for mutations in the MAGT1 gene being associated with congenital disorders of glycosylation. Mutations in the MAGT1 gene are associated with a form of X-linked intellectual disability (XLID).
The MAGT1 gene is located on the X chromosome (Xq21.1) and is composed of 10 exons that generate two alternatively spliced mRNAs. The primary MAGT1 encoded protein is 367 amino acids. The gene has multiple in-frame translational initiation sites, therefore, allowing for different proteins to be translated. This magnesium transporter is a member of the solute carrier family of transporter and as such is also identified as SLC58A1.
TUSC3-CDG is an autosomal recessive disorder that results from deficiencies in the tumor suppressor candidate 3 gene (symbol: TUSC3). The TUSC3 encoded protein interacts with the N-oligosaccharyltransferase complex (OST) which is the enzyme complex that transfers the lipid-linked oligosaccharide (LLO) structure (Glc3Man9GlcNAc2-Dol) to the asparagine residue in nascent polypeptides. This reaction is the central step in N-linked glycosylation of proteins. Mutations in the TUCS3 gene are associated with autosomal recessive nonsyndromic mental retardation-7 (MRT7). At least 13 patients have been identified harboring mutations in this gene and all suffer from severe nonsyndromic intellectual impairment. The phenotype observed in these patients and the identification of the product of the TUSC3 gene being involved in N-glycosylation implicates a role for this process in higher brain functions.
The TUSC3 gene is located on chromosome 8p22 and is composed of 13 exons that generate three alternatively spliced mRNAs that encode two distinct protein isoforms, one a 348 amino acid precursor identified as isoform a and the other a 347 amino acid precursor identified as isoform b. The TUSC3 encoded protein is also known to function as a magnesium transporter of the SLC family and is, therefore, also identified as SLC58A2.
Type II CDG Forms
CDG-IIa (MGAT2-CDG; CDG2A) is an autosomal recessive disorder that results from loss of the enzyme α-1,6-mannosyl-glycoprotein β-1,2-N-acetylglucosaminyltransferase (GlcNAc-TII; also called UDP-N-acetylglucosamine:α-6-D-mannoside-β-1,2-N-acetylglucosaminyltransferase II) which is encoded by the MGAT2 gene. The results of this defect are significant loss of complex sialylated structures. Following transfer of the en bloc oligosaccharide to Asn residues, only monosialylated structures are made. Symptoms of CDG-IIa include severe psychomotor impairment, hypotonia, coarse facies and frequent infections. In experiments in mice it was found that MGAT2 null animals all died within 4 weeks after birth and a significant number die in utero from gastrointestinal blockage. From these results it is speculated that the true incidence of human MGAT2 defects may go undetected due to spontaneous fetal abortion or death shortly after birth.
The MGAT2 gene is located on chromosome 14q21.3 and is an intronless gene encoding a protein of 447 amino acids.
CDG-IIb (GCS1-CDG; CDG2B) is an autosomal recessive disorder that results from deficiencies in mannosyl-oligosaccharide glucosidase 1 (gene symbol: MOGS; also called glucosidase 1: GCS1). A patient identified with CDG-IIb exhibited dysmorphic features, seizures, hypotonia, hepatomegaly, feeding difficulty, and failure to thrive. This patient did not survive. To date a total of 24 individuals have been identified with GCS1-CDG. The common spectrum of pathologies observed in these patients includes facial dysmorphisms, global developmental delay, muscular hypotonia, seizures, immune dysfunction related to hypogammaglobulinemia, and hepatomegaly. A relevant clinical finding in all GCS1-CDG patients is elevation in the tetrasaccharide, Glc3-Man, in the urine.
The MOGS gene is located on chromosome 2p13.1 and is composed of 5 exons that generate two alternatively spliced mRNAs encoding MOGS isoform 1 (837 amino acids) and MOGS isoform 2 (731 amino acids).
CDG-IIc (SLC35C1-CDG; CDG2C) is an autosomal recessive disorder that is more commonly referred to as leukocyte adhesion deficiency syndrome II (LAD II). LAD II belongs to the class of disorders referred to as primary immunodeficiency syndromes as the symptoms of the disease manifest due to defects in leukocyte function. Symptoms of LAD II are characterized by unique facial features, recurrent infections, persistent leukocytosis, defective neutrophil chemotaxis and severe growth and intellectual impairment. The genetic defect resulting in LAD II is in the pathway of fucose utilization leading to loss of fucosylated glycans on the cell surface. An additional feature of LAD II is that individuals harbor the rare Bombay (hh) blood type at the ABO locus as well as lack the Lewis blood group antigens. The Bombay blood type is characterized by a deficiency in the H (referred to as the O-type), A and B antigens due to loss of the fucose residue. Each of these blood group antigens contains a Fuc-α-1,2-Gal modification that is the final carbohydrate addition to these antigens. These fucosylation reactions are catalyzed by α-1,2-fucosyltransferase which is encoded by the H and Se loci. The defective neutrophil chemotaxis is due to the loss of a selectin ligand on these cells. This ligand is the sialylated Lewisx antigen, another blood group antigen.
The recurrent infections seen in LAD II patients are the result of the defective neutrophil function. Neutrophils are involved in innate immunity responses to bacterial infection. To carry out their role in host defense mechanisms, neutrophils must adhere to the surface of the endothelium at the site of inflammation which is an event mediated by cell surface adhesion molecules. The selectin family (E-, L-, and P-selectins) of animal lectins are necessary to mediate the initial process of neutrophil adherence to the endothelium. The selectins recognize sialylated fucosylated lactosamines typified by the Lewisx antigen. Once neutrophils adhere and roll along the surface of the endothelium (due to vascular flow), the integrin family of adhesion molecules allow for firm adherence followed by tissue penetration. A related disorder, termed LAD I, is caused by the absence of CD18 which is the β2 subunit of the leukocyte integrin found on the surface of neutrophils and monocytes.
Because there is widespread loss of fucosylated antigens in LAD II patients, each of which can be formed through the actions of several fucosyltransferases, the role of these enzymes in the disease could be ruled out. In addition, normal levels of the α-1,2-, α-1,3- and α-1,4-fucosyltransferases were observed in the serum of LAD II individuals. These observations indicated that the pathways to GDP-fucose synthesis or utilization by the fucosyltransferases in the Golgi must be deficient in LAD II. Fucose can be converted to GDP-fucose by salvage of free fucose (exogenous or derived through glycoconjugate degradation) or by epimerization of GDP-mannose. In order for GDP-fucose to be utilized by Golgi fucosyltransferases it must first be transported into the Golgi from the cytosol where it is synthesized. Examinations of the enzymes involved in the synthesis and transport of GDP-fucose have been undertaken. While the activity of one of the enzymes of GDP-mannose epimerization (GDP-D-mannose 4,6-dehydratase, GMD) has been shown to be reduced in LAD II patients, it has been determined that the major defect causing LAD II is an impairment in the transport of GDP-fucose into the Golgi. This latter reaction is catalyzed by the GDP-fucose transporter encoded by the FUCT1 gene (also identified as solute carrier family 35, member C1: SCL35C1).
The SLC35C1 gene is located on chromosome 11p11.2 and is composed of 4 exons that generate three alternatively spliced mRNAs that encode two different protein isoforms. Isoform a is 364 amino acids and isoform b is 351 amino acids.
CDG-IId (B4GALT1-CDG; CDG2D) is an autosomal recessive disorder that results from deficiencies in UDP-Gal:β-GlcNAc β-1,4- galactosyltransferase, polypeptide 1 (gene symbol: B4GALT1). There are seven B4GALT genes in the human genome. All of the encoded enzymes are type II membrane-bound glycoproteins that exhibit exclusive specificity for UDP-galactose as a donor substrate that is transferred to either Glc, GlcNAc, or xylulose (Xyl). The enzyme encoded by the B4GALT1 gene is unique among the seven B4GALT genes in that, dependent upon the tissue of expression, it exhibits distinct activities. The major B4GALT1 encoded activity adds galactose to N-acetylglucosamine residues that are either monosaccharides or the nonreducing ends of carbohydrate chains in glycoprotein. This enzyme activity is encoded by the longer of two alternative mRNAs that are generated from the B4GALT1 gene through the use of alternative transcriptional initiation sites. The shorter mRNA is restricted to lactating mammary tissue and, although the encoded protein is identical to that encoded by the longer mRNA, it undergoes distinct post-translational modification and forms a heterodimer with lactalbumin to catalyze the synthesis of the disaccharide, lactose. Only one patient has been identified with CDG-IId. The child exhibited psychomotor impairment, severe spontaneous bleeding, hydrocephalus, and myopathy.
The B4GALT1 gene is located on chromosome 9p21.1 and is composed of 11 exons that generate the four mRNAs through alternative transcriptional initiation sites and alternative mRNA splicing.
CDG-IIe (COG7-CDG; CDG2E) is an autosomal recessive disorder that results from deficiencies in the gene encoding component of oligomeric Golgi complex-7 (gene symbol: COG7). The protein encoded by the COG7 gene is one of eight subunits of the conserved oligomeric Golgi (COG) complex. Because this gene defect disrupts proper Golgi trafficking its effects are evident in both the processes of N– and O-linked glycosylation pathways. Patients present with growth impairment, progressive severe microcephaly, hypotonia, adducted thumbs, feeding problems due to gastrointestinal pseudoobstruction, failure to thrive, cardiac anomalies, wrinkled skin, and episodes of extreme hyperthermia.
The COG7 gene is located on chromosome 16p12.2 and is composed of 18 exons that encode a protein of 770 amino acids.
CDG-IIf (SLC35A1-CDG; CDG2F) is an autosomal recessive disorder that results from deficiencies in the nucleotide sugar transporter (NST) responsible for the transport of CMP-sialic acid (also called CMP-NANA) from the cytosol into the lumen of the ER and/or Golgi apparatus. This transporter is a member of the solute carrier family of transporters (gene symbol: SLC35A1). Deficiencies in this transporter result in marked thrombocytopenia and neutropenia as well as recurrent infections. The recurrent infections are attributed to undetectable sialylated Lewisx on the surface of neutrophils which prevents them from adhering to the endothelium at sites of inflammation.
The SLC35A1 gene is located on chromosome 6q15 and is composed of 8 exons that generate two alternatively spliced mRNAs encoding SLC35A1 isoform a (337 amino acids) and isoform b (278 amino acids).
CDG-IIg (COG1-CDG; CDG2G) is an autosomal recessive disorder that results from deficiencies in the gene encoding component of oligomeric Golgi complex-1 (gene symbol: COG1). The protein encoded by the COG1 gene is one of eight subunits of the conserved oligomeric Golgi (COG) complex. Because this gene defect disrupts proper Golgi trafficking its effects are evident in both the processes of N– and O-linked glycosylation pathways. Few cases of CDG-IIg have been described but all have presented in infancy with failure to thrive, generalized hypotonia, growth impairment with rhizomelic short stature, and mild psychomotor impairment. In one patient the disease was described as a cerebrocostomandibular (CCMS)-like syndrome.
The COG1 gene is located on chromosome 17q25.1 and is composed of 14 exons that encode a protein of 980 amino acids.
CDG-IIh (COG8-CDG; CDG2H) results from deficiencies in the gene encoding component of oligomeric Golgi complex-8 (gene symbol: COG8). The precise mode of inheritance of CDG-IIh is not known but is most likely autosomal recessive. The protein encoded by the COG8 gene is one of eight subunits of the conserved oligomeric Golgi (COG) complex. Because this gene defect disrupts proper Golgi trafficking its effects are evident in both the processes of N– and O-linked glycosylation pathways. Only two cases of CDG-IIh have been described and both patients suffered from severe psychomotor impairment, failure to thrive, seizures, and dairy and wheat product intolerance.
The COG8 gene is located on chromosome 16q22.1 and is composed of 7 exons that generate eight alternatively spliced mRNAs, that collectively encode seven distinct protein isoforms.
CDG-IIi (COG5-CDG; CDG2I) is an autosomal recessive disorder that results from deficiencies in the gene encoding component of oligomeric Golgi complex-5 (gene symbol: COG5). The protein encoded by the COG5 gene is one of eight subunits of the conserved oligomeric Golgi (COG) complex. Because this gene defect disrupts proper Golgi trafficking its effects are evident in both the processes of N– and O-linked glycosylation pathways. A single case of CDG-IIi has been described where the patient suffered from moderate intellectual impairment with slow and inarticulate speech, truncal ataxia, and mild hypotonia.
The COG5 gene is located on chromosome 7q22.3 and is composed of 23 exons that generate nine alternatively spliced mRNAs, each of which encode a distinct protein isoform.
CDG-IIj (COG4-CDG; CDG2J) is an autosomal recessive disorder that results from deficiencies in the gene encoding component of oligomeric Golgi complex-4 (gene symbol: COG4). The protein encoded by the COG4 gene is one of eight subunits of the conserved oligomeric Golgi (COG) complex. Because this gene defect disrupts proper Golgi trafficking its effects are evident in both the processes of N– and O-linked glycosylation pathways. Two cases of CDG-IIj have been described where one patient suffered from milder symptoms including dysmorphic features, such as down-sloping frontal area and thick hair, mild neurologic signs, including axial hypotonia, mild peripheral hypertonia, and hyperreflexia. The other case involved a child with profound developmental delay, hypotonia, nystagmus, hepatosplenomegaly, and poor growth ultimately resulting in seizures, liver cirrhosis, and recurrent infections that were ultimately fatal around age 2 years.
The COG4 gene is located on chromosome 16q22.1 (close to the location of the COG8 gene) and is composed of 20 exons that generate three alternatively spliced mRNAs encoding three distinct protein isoforms: isoform 1 (789 amino acids), isoform 2 (768 amino acids), and isoform 3 (647 amino acids).
CDG-IIk (TMEM165-CDG; CDG2K) is an autosomal recessive disorder that results from deficiencies in transmembrane protein 165 (gene symbol: TMEM165) which plays a role in protein sialylation in the Golgi apparatus. Patients with CDG-IIk exhibit psychomotor and growth impairment, short stature, dysmorphism, hypotonia, eye abnormalities, acquired microcephaly, hepatomegaly, and skeletal dysplasia.
The TMEM165 gene is located on chromosome 4q12 and is composed of 9 exons that generate two alternatively spliced mRNAs, one of which encodes a precursor protein of 324 amino acids while the other has a frameshift resulting in an early stop codon, thus producing a non-functional protein. This second mRNA is targeted for nonsense-mediated mRNA decay (NMD).
CDG-IIL (COG6-CDG; CDG2L) is an autosomal recessive disorder that results from deficiencies in the gene encoding component of oligomeric Golgi complex-6 (gene symbol: COG6). The protein encoded by the COG6 gene is one of eight subunits of the conserved oligomeric Golgi (COG) complex. Because this gene defect disrupts proper Golgi trafficking its effects are evident in both the processes of N– and O-linked glycosylation pathways. Few cases of CDG-IIL has been described but the severity of the disorder is profound resulting in early lethality. Symptoms observed in afflicted infants include intractable focal seizures, dysmorphic features, including microcephaly, postaxial polydactyly, broad palpebral fissures, retrognathia, recurrent infections, diarrhea, and primary combined immunodeficiency.
The COG6 gene is located on chromosome 13q14.11 and is composed of 22 exons that generate two alternatively spliced mRNAs encoding two distinct protein isoforms: isoform 1 (657 amino acids) and isoform 2 (615 amino acids).
CDG-IIm (SLC35A2-CDG; CDG2M) is an X-lined dominant disorder that results from deficiencies in the solute carrier transporter family 35 member A2 (gene symbol: SLC35A2) which is a nucleotide-sugar transporter that transports UDP-galactose from the cytosol into Golgi vesicles. Patients with CDG-IIm exhibit early-infantile epileptic encephalopathy, developmental delay, hypotonia, variable ocular anomalies, seizures, hypsarrhythmia, poor feeding, microcephaly, recurrent infections, dysmorphic features, shortened limbs, and coagulation defects. Some patients with CDG-IIm have been shown to be the result of somatic mosaicism and not due to parental transmission of a dominant X-linked mutation.
The SLC35A2 gene is located on the X chromosome (Xp11.23) and is composed of 6 exons that generate eight alternatively spliced mRNAs encoding eight distinct protein isoforms.
CDG-IIn (SLC39A8-CDG; CDG2N) is an autosomal recessive disorder that results from deficiencies in the solute carrier transporter family 39 member A8 (gene symbol: SLC39A8) which found in the plasma membrane and mitochondria where it functions in the cellular import of zinc at the onset of inflammation. Several of the patients described with CDG-IIn were of Hutterite descent. The symptoms observed in CDG-IIn include profound psychomotor impairment with delayed head control, severe hypotonia, inability to walk, variable ability to sit independently, and profound intellectual disability, strabismus, recurrent infections, and seizures.
The SLC39A8 gene is located on chromosome 4q24 and is composed of 17 exons that generate four alternatively spliced mRNAs encoding three distinct protein isoforms.
CDG-IIo (CCDC115-CDG; CDG2O) is an autosomal recessive disorder that results from deficiencies in coiled-coil domain containing 115 protein (gene symbol: CCDC115) which localizes to the endoplasmic reticulum-Golgi intermediate compartment (ERGIC). Patients with CDG-IIo exhibit infantile onset of progressive liver failure, hypotonia, and delayed psychomotor development resulting in early lethality.
The CCDC115 gene is located on chromosome 2q21.1 and is composed of 6 exons that generate three alternatively spliced mRNAs encoding three distinct protein isoforms.
CDG-IIp (TMEM199-CDG; CDG2P) is an autosomal recessive disorder that is an autosomal recessive disorder that results from deficiencies in transmembrane protein 199 (gene symbol: TMEM199) which localizes to the endoplasmic reticulum-Golgi intermediate compartment (ERGIC) and coat protein complex I (COPI). Patients with CDG-IIp primarily manifest with a mild metabolic disorder primarily affecting the liver that presents in early adolescence.
The TMEM199 gene is located on chromosome 17q11.2 and is composed of 6 exons that encode a protein of 208 amino acids.
Protein O-glycosylation Defects:
MDDG: Several disorders are known that result from defects in the O-mannosylation of the α-dystroglycan protein of the dystrophin-glycoprotein complex (DGC) and are referred to as muscular dystrophy-dystroglycanopathies (MDDG). There are at least six related syndromes identified as MDDG (MDDGA1, MDDGA2, MDDGA3, MDDGA4 (and MDDGB4), MDDGA5 (and MDDGB5), and MDDGA6 (and MDDGB6). The letter “A” and “B” in the designations relate to disease severity with “A” indicating severe, and “B” indicating intermediate.
Several related disorders that result from defects in O-mannosylation were originally classified as Walker-Warburg syndrome (WWS; most severe forms) and the muscle-eye-brain diseases (MEB; less severe forms). These disorders result from deficiencies in either POMT1, POMT2, or POMGNT1. The POMT1 and POMT2 genes encode O-mannosyltransferase-1 and -2, respectively. The POMGNT1 gene encodes O-mannose β-1,2-N-acetylglucosaminyltransferase.
The proposed nomenclature for CDG that cause congenital muscular dystrophy spectrum due to defects in POMT1 (POMT1-CDG), POMT2 (POMT2-CDG), and POMGNT1 (POMGNT1-CDG) is MDDGA1, MDDGA2, and MDDGA3, respectively. These three MDDG disorders are associated with characteristic brain and eye malformations, profound intellectual impairment, congenital muscular dystrophy, and early death.
MDDGA4 is also commonly referred to as Fukuyama-type congenital muscular dystrophy and is caused by mutations in the fukutin gene (symbol: FKTN) whose encoded protein is thought to be a glycosyltransferase involved in brain development.
MDDGA5 (also referred to as congenital muscular dystrophy type 1C) results from mutations in the fukutin-related protein gene (FKRP).
MDDGA6 (also referred to as congenital muscular dystrophy type 1D) results from mutations in the LARGE1 (like-acetylglucosaminyltransferase 1) gene which encodes the bifunctional enzyme identified as LARGE xylulose- and glucuronyltransferase 1. The LARGE1 encoded enzyme belongs to the N-acetylglucosaminyltransferase family of enzymes. The LARGE1 encoded enzyme is important in the glycosylation of α-dystroglycan.
B4GALT7-CDG: Multiple cartilaginous exostoses (MCE) and a progeroid variant (accelerated aging) of Ehlers-Danlos syndrome, EDS (EDS with short stature and limb anomalies, EDSSLA) are the result of defects in O-xylosylation. Additional forms of MCE result from defects in either the EXT1 or EXT2 genes which encode glycosyltransferases involved in heparin and heparan sulfate biosynthesis.
The EDSSLA disorder is caused by a protein O-glycosylation defect that is the result of deficiency in the B4GALT7 gene encoding β-1,4-galactosyltransferase 7. This gene is also identified by the name xylosylprotein 4-β-galactosyltransferase (XGALT1 or XGPT1). The proposed CDG nomenclature for the EDSSLA is B4GALT7-CDG.
GALNT3-CDG: Hyperphosphatemic familial tumoral calcinosis (HFTC) results from defects in at least three genes. One form of HFTC results from a defect in O–N-acetylgalactosaminylation which is catalyzed by the GALNT3 gene encoding polypeptide N-acetylgalactosaminyltransferase 3. The proposed nomenclature for the GALNT3 deficient CDG is GALNT3-CDG.
Additional causes of HFTC are not related to protein glycosylation but involve genes encoding participants in signal transduction by fibroblast growth factors (FGF), specifically mutations in FGF23 (gene symbol: FGF23) and KLOTHO (gene symbol: KL). FGF23 is known to interact with KLOTHO and then the complex binds to its receptor which is predominantly FGF receptor 1 (FGFR1). The KLOTHO gene was originally isolated from a mouse model of age-related disorders and thus the gene was named after the Fate of Greek mythology who spins the thread of life.
LFNG-CDG: Spondylocostal dysostosis type 3 results from a defect in O-fucosylation of members of the Notch receptor family. This disorder is associated with vertebral segmentation problems and rib anomalies. The fucosylation reaction is catalyzed by O-fucose-specific β-1,3-N-acetylglucosaminyltransferase encoded by the lunatic fringe (LFNG) gene. The lunatic fringe protein is the human homolog of the fruit fly gene fringe which was identified as a mutation resulting in defective early development.
Lunatic fringe modulates the activation of the Notch receptor signal transduction pathway. Lunatic fringe incorporates O-fucose residues into the epidermal growth factor-like (EGF-like) repeats of Notch receptors of which there are four (NOTCH1–NOTCH4) in humans.